Yeast DSB Repair
Overview
The longest continuous research effort in the Wilson group is the study of DNA double-strand break (DSB) repair mechanisms, mainly using the yeast Saccharomyces cerevisiae as a model genetic organism. DSBs are a serious genomic lesion that directly threatens the formation of chromosomal rearrangements.
Cells have (at least) two DSB repair mechanisms to restore genomic integrity: nonhomologous end joining (NHEJ), i.e., the direct religation of DSB ends, and homologous recombination (HR), which copies an intact “donor” locus to restore integrity at the broken allele. Our work has historically focused on NHEJ as an error-prone mechanism that is often implicated in the formation of de novo structural variant (SV) junctions. However, we also study the early steps of HR, namely, 5’ resection of DSB ends, as it is the primary alternative to NHEJ. Like many, we think that proper control of the decision point between NHEJ and HR is a key factor in determining the accuracy of DSB repair outcomes.
DNA double-strand break (DSB) repair pathways. DSB repair is best thought of as an integrated set of processes in which coordinated, sequential actions help to optimize repair accuracy in different cellular contexts.
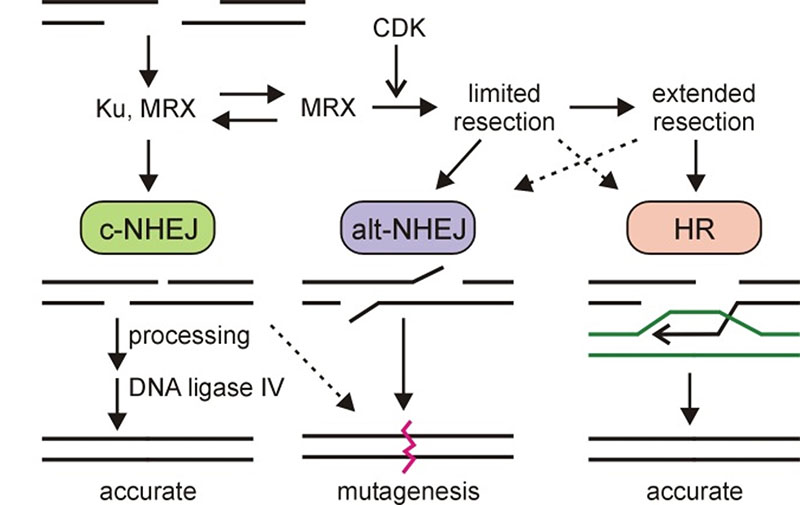
DNA double-strand break (DSB) repair pathways. DSB repair is best thought of as an integrated set of processes in which coordinated, sequential actions help to optimize repair accuracy in different cellular contexts.
Goals and directions
We put our work on yeast DSB repair into the broader context of Dr. Wilson’s efforts. We use yeast as a model that allows certain classes of experiments to be performed in vivo that are still not possible in mammalian cells. Our other project areas show how an understanding of basic DSB repair mechanisms developed in reductionist systems can directly impact studies of mammalian mutagenesis.
Current focus areas in the yeast lab include:
- understanding how spatial factors in the nucleus influence chromosomal rearrangement by NHEJ
- continuing work to understand the precise biochemical events that coordinate NHEJ vs. HR execution in living cells